Comparative Population Genetics in the Human Gut Microbiome
We are delighted to share our latest preprint on:
Comparative Population Genetics in the Human Gut Microbiome
William Shoemaker, Daisy Chen, and Nandita Garud
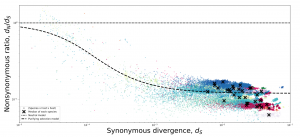
The genetic variation in the human gut microbiome is responsible for conferring a number of crucial phenotypes like the ability to digest food and metabolize drugs. Yet, our understanding of how this variation arises and is maintained remains relatively poor. Thus, the microbiome remains a largely untapped resource, as the large number of co-existing species in this microbiome presents a unique opportunity to compare and contrast evolutionary processes across species to identify universal trends and deviations. Here we outline features of the human gut microbiome that, while not unique in isolation, as an assemblage make it a system with unparalleled potential for comparative population genomics studies. We consciously take a broad view of comparative population genetics, emphasizing how sampling a large number of species allows researchers to identify universal evolutionary dynamics in addition to new genes, which can then be leveraged to identify exceptional species that deviate from general patterns. To highlight the potential power of comparative population genetics in the microbiome, we re-analyzed patterns of purifying selection across ~40 prevalent species in the human gut microbiome to identify intriguing trends which highlight functional categories in the microbiome that may be under more or less constraint.